The BioCatalysis team was pioneer in the functional characterization of Radical-SAM enzymes with the Anaerobic Ribonucleotide Reductase activating enzyme (RNR-AE)
[Reference], one of the prototypes of this class of enzymes in addition with the Lysine Aminomutase (LAM) and the Pyruvate Formate Lyase activating enzyme (PFL-AE)1. From then on, the Biocatalysis team studied some others (Biotin Synthase, Lipoate Synthase, Spore photoproduct Lyase, MiaB, RIMO, …). Radical-SAM enzyme catalyze a large amount of reactions and currently, our researches are focusing on Radical-SAM enzymes involved in post-transcriptional tRNA modifications and in post-translational peptide modification.
Post-transcriptional tRNA modifications
M. Atta, V. Duarte, P. Carpentier
Since many years, the BioCatalysis team is interested in the enzymology of post-transcriptional modification of tRNA. In fact, the enzymes that generate the modified nucleosides are appealing targets for studies for several reasons, as they offer insight into protein-RNA interactions. They must discriminate between substrate and non-substrate RNA as well as substrate and product nucleosides within the same RNA context, i
.e. they must discriminate between multiple tRNAs of globally identical shape. Furthermore, the RNA-modifying enzymes conduct many chemically sophisticated reactions that merit examination in their own right. The favorite biosynthetic pathway studied in the team is the formation of 2-methylthio-N-6-(cis-hydroxy)isopentenyl adenosine and it’s derivative (ms
2io
6A
37/ms
2ms
2i
6A
37/ ms
2ms
2io
6A
37) which is present at position 37 adjacent to the anticodon of specific tRNAs (Figure 1). Among the enzymes involved in this process (MiaA, MiaB, MiaE), two are iron-containing enzymes. MiaB belongs to the radical-SAM enzyme family ([4Fe-4S]-containing enzyme) and MiaE is a binuclear iron-containing enzyme. Both involve a C-H bond activation using a radical mechanism, allowing methylthiol insertion in the case of MiaB and an hydroxyl group insertion for MiaE. While MiaB is oxygen sensitive, MiaE needs dioxygen. Currently, the objective is to understand at a very fundamental level, by combining the methods of microbial genetics, protein chemistry, and advanced spectroscopy, the mechanistic and physiological features of the sophisticated radical-based chemistry that installed sulfur and oxygen atoms into biological tRNA molecule (Figure 1).
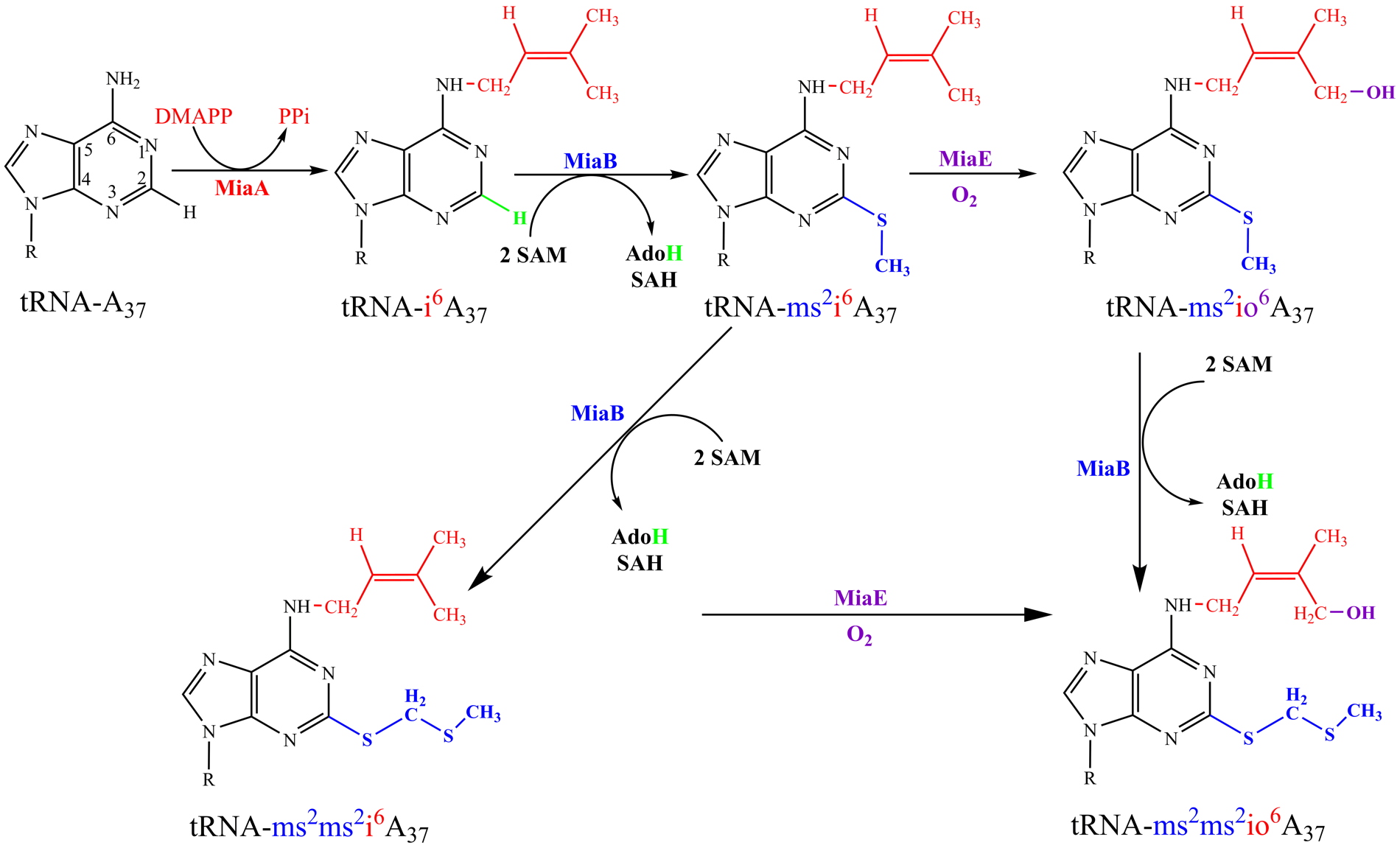
Figure 1: (A) Established biosynthetic pathway of ms
2io
6A
37. The enzymes involved in this pathway are: MiaA, MiaB, and MiaE. DMAPP, dimethylallyl diphosphate; PPi, pyrophosphate; SAM, S-adenosylmethionine; SAH, S-adenosylhomocysteine; AdoH, 5’-deoxyadenosyl. (B) The newly discovered biosynthesis of a thioacetal ms
2ms
2i
6A
37. (C) A hypothetical hydroxylation of thioacetal by MiaE enzyme.
 Selected publications
Dal Magro C, Keller P, Kotter A, Werner S, Duarte V, Marchand V, Ignarski M, Freiwald A, Muller RU, Dieterich C, Motorin Y, Butter F, Atta M and Helm M
Selected publications
Dal Magro C, Keller P, Kotter A, Werner S, Duarte V, Marchand V, Ignarski M, Freiwald A, Muller RU, Dieterich C, Motorin Y, Butter F, Atta M and Helm M
A vastly increased chemical variety of RNA modifications containing a thioacetal structure.
Angewandte Chemie-International Edition, 2018,
57(26): 7893-7897
Arragain S, Bimai O, Legrand P, Caillat S, Ravanat JL, Touati N, Binet L, Atta M, Fontecave M, Golinelli-Pimpaneau B and Buckel W Nonredox thiolation in tRNA occurring
via sulfur activation by a [4Fe-4S] cluster.
Proceedings of the National Academy of Sciences of the USA, 2017,
114(28): 7355-7360
Mulliez E, Duarte V, Arragain S, Fontecave M and Atta M On the role of additional 4 Fe-4S clusters with a free coordination site in radical-SAM enzymes.
Frontiers in Chemistry, 2017,
5: Article Number 17
Forouhar F, Arragain S, Atta M, Gambarelli S, Mouesca JM, Hussain M, Xiao R, Kieffer-Jaquinod S, Seetharaman J, Acton TB, Montelione GT, Mulliez E, Hunt JF and Fontecave M Two Fe-S clusters catalyze sulfur insertion by radical-SAM methylthiotransferases.
Nature Chemical Biology, 2013,
9(5): 333-338
Mathevon C, Pierrel F, Oddou JL, Garcia-Serres R, Blondin G, Latour JM, Menage S, Gambarelli S, Fontecave M and Atta M tRNA-modifying MiaE protein from
Salmonella typhimurium is a nonheme diiron monooxygenase.
Proceedings of the National Academy of Sciences USA, 2007,
104(33): 13295-13300
Post-translational peptide modification: Biosynthesis of thioether-bond
V. Duarte, M. Atta, C. Basset, L. Shamseddine (PhD CFR CEA 2020-2023)
Few years ago, BioCatalysis team characterized the RimO Radical-SAM enzyme (MiaB-like enzyme) involved in the methyl-thiolation of the
E. coli ribosomal protein S12. Currently, the team focuses on Radical-SAM enzymes involved in the modification of RiPPS (Ribosomally synthesized and Post-translationally modified Peptides). These Radical SAM enzymes catalyze a wide range of post-translational modifications including methylation, thioether and carbon-carbon bond formation, complex rearrangements and epimerization. Our work concerns the thioether cross-links between cysteine sulfur and either the α
¯, ß- or the γ- carbon of a partner amino acid (Figure 2). This mechanistic difference leads to two classes of RiPPs, the Sactipeptides (C-α thioether) and the Ranthipeptides (C-ß or the C-γ thiother).
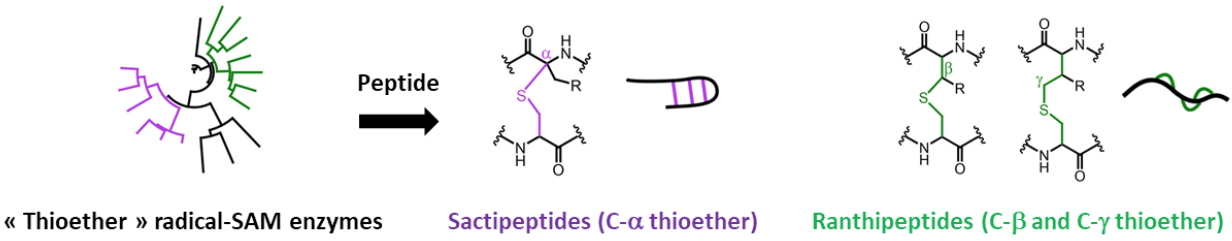
Figure 2: Radical SAM enzymes and the biosynthesis of sactipeptides and ranthipeptides.
The enzymes within the radical SAM superfamily contain the canonical CysX
3CysX
2Cys motif, that binds the radical-SAM [4Fe-4S]
2+/1+ cluster, in which the fourth, unique iron, is used to bind the SAM cofactor. In its reduced state, the [4Fe-4S]
1+ cluster catalyzes the reductive cleavage of SAM to generate a 5′-deoxyadenosyl radical (5’-Ado
•). This radical abstracts an H atom from the cognate substrate to initiate catalysis (Figure 3).

Figure 3: Reductive cleavage of S-Adenosylmethionine by radical SAM enzymes.
Structural and functional data indicate that radical SAM enzymes involved in the sactipeptides or ranthipeptides biosynthesis contain a C-terminal extension appended to the radical-SAM cluster called SPASM or Twitch domains that house additional [4Fe-4S]
2+ clusters. In contrast to the Twitch domains, which bind only one additional cluster, SPASM domains present a conserved cysteine-rich motif that coordinates two additional iron-sulfur clusters. The role of these clusters waits for clarification, but it is suggested that they possibly interact with the substrate during catalysis or may also be implicated in electrons transfer. Our research work investigates novel radical SAM enzymes involved in the biosynthesis of thioether bond containing antimicrobial peptides. The goal is to better understand how these enzymes install the post-translational modifications in a selective way by: i) deciphering the role of each Fe-S cluster of the proteins; ii) understanding how the enzymes interact with the substrate; iii) investigating the role of the leader sequence present on the N-terminal part of the precursor peptides. Overall, this work will allow us to explore the structural and functional diversities of the radical SAM enzymes and their corresponding substrates.
The BioCatalysis team is interested by the RiPPs biosynthesis from a basic point of view but also from a health point of view, as
RiPPs constitute promising antimicrobial agents for a potential therapeutic use.